

Research
The research at IBP bridges between physical mechanisms in living cells and the evolutionary dynamics in populations.

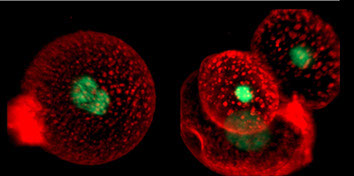
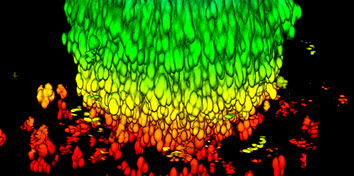
We work on inverse statistical problems and stochastic systems in order to understand information processing in biological systems and biological evolution. Recent applications are in the fields of drug resistance to cancer therapy, where we do both theoretical modelling and cell line experiments, the inference of signalling networks, and stochastic dynamics in gene regulation.
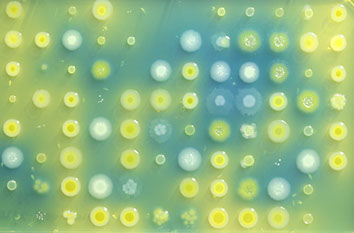

How do cells respond to combinations of drugs and other signals? Can the evolution of antibiotic resistance be predicted and prevented? How do microbial ecosystems function? To tackle questions like these and ultimately discover the general principles and laws of living systems, we combine concepts and techniques from physics with molecular biology and advanced high-throughput experiments.
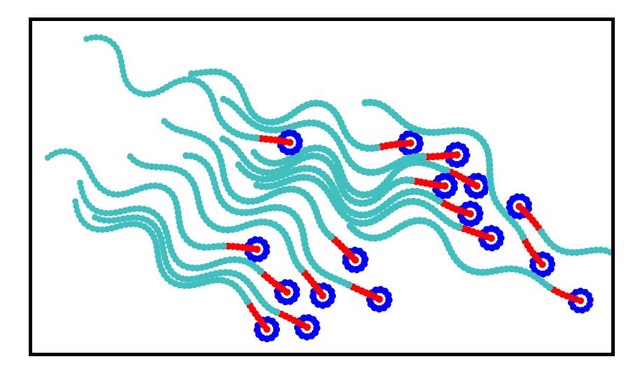
We develop theoretical and numerical models to understand physical mechanisms governing the structure and dynamics of soft, active, and living matter. Research topics include the interactions of proteins and nanoparticles with membranes, the behaviour of blood cells under flow, the non-equilibrium dynamics of the cytoskeleton, the motion and flow generation of microswimmers (bacteria, sperm, cilia, artificial self-propelled particles), and the growth of tissues. Our overarching goal is to elucidate the physical principles of non-equilibrium -- active and driven — systems.
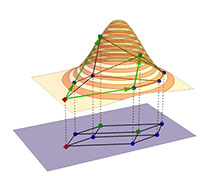
We develop population genetic models to describe and analyse how populations evolve to adapt to a new environment. A key goal is to understand the factors that shape the pathways along which evolution can proceed and thus ultimately determine the predictability of the evolutionary process. Applications derive mostly from the field of experimental evolution, in particular the evolution of drug resistance in microbes.
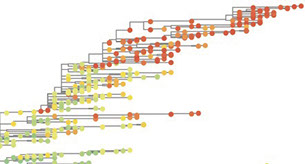
Our lab works on living systems and their evolution. How does this dynamics work? Modern molecular biological data provide unprecedented opportunities to understand evolution in a quantitative, empirical way. Conversely, evolutionary analysis becomes a tool to understand biological function. We work with data from fast-evolving natural populations and from laboratory evolution experiments, including the influenza virus, E. coli, and yeast. Our goal is to make evolutionary biology a predictive science.
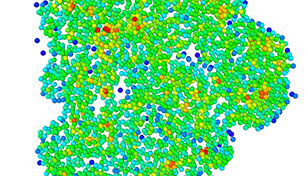
How do bacteria control molecular forces to shape multicellular communities called biofilms? Can we correlate biofilm structure with tolerance against drug treatment? These questions are at the center of research at the single cell level. At the population level, we design methods for quantifying costs and benefits of horizontal gene transfer between bacteria. In our lab, physicists and biologists work in close collaboration using tools from nanotechnology, image analysis, molecular biology, and experimental evolution.
Research Centers
Collaborative Research Center 1310 Predictability in Evolution
This DFG-funded Center (scientific coordinator: Michael Lässig, IBP) focuses on predictive analysis of fast-evolving systems: microbes, viruses, cancer, and immune systems. It links all research groups of IBP with Computational Biology and Medicine at UoC, as well as other national and international institutions (Heinrich-Heine-University Düsseldorf, ENS Paris, Wageningen University, MPI for Dynamics and Self-Organization Göttingen, IGC Lisbon).
DFG Priority Program Probabilistic Structures in Evolution
A Germany-wide consortium devoted to the in-depth theoretical study of stochastic processes in population genetics.
UoC Center for Data and Simulation Science
This Center is UoC’s over-arching structure to link natural sciences, life sciences, economics, social sciences, and humanities with core competence in data science and scientific computing.
UoC Forum Classical and Quantum Dynamics of Interacting Particle Systems
The Forum brings together researchers from probability theory and theoretical physics in the Cologne-Bonn area with an interest in a broad range of aspects of interacting particle systems.
imprint

